Research in the Teitell group focuses on mechanisms of cancer formation and progression. We place a special emphasis on studies of leukemia and lymphoma that arise during normal B cell development. Our work has evolved into several areas with distinct sets of techniques and approaches; however, all project areas address fundamental questions concerning the etiology and progression of cancer, providing a common link for the group. Areas of active research in our group include:
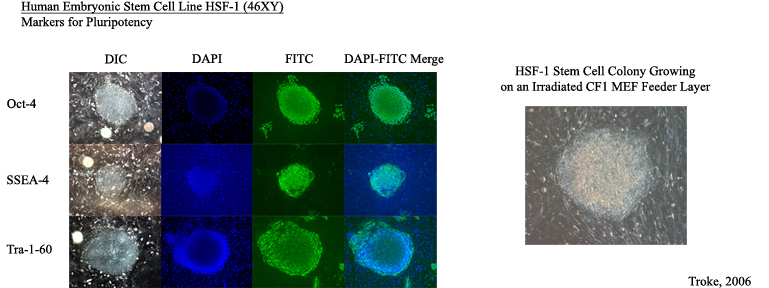
Our group discovered a role for aberrant TCL1 oncogene expression in human leukemias and lymphomas and generated a proof-of-principle TCL1 transgenic mouse model that develops mainly germinal center (GC) B cell cancers (Teitell PNAS 1999; Hoyer PNAS 2002; Teitell Nature Reviews Cancer 2005). Studies of TCL1 gene regulation in B cell development and cancer ( PNAS 2007) led to the identification of a signal transduction pathway that controls the terminal differentiation of GC B cells into antibody-secreting plasma cells (Sherman Molecular Cell 2010; Sherman Trends in Cell Biology 2011). Key steps in this signal transduction pathway include the activation of the tumor suppressor LKB1 and that of an unknown AMPK (5’ AMP activated protein kinase) family member protein, which results in the nuclear export and inactivation of the CREB transcriptional co-activator, CRTC2 (figure). Inactivation of CRTC2 represses direct target genes to enable B cells to exit the GC reaction and differentiate into plasma cells. Failure to inactivate CRTC2 blocks differentiation and locks down GC B cells in a state of continued growth and increased resistance to apoptosis, thereby generating an environment that fosters tumor formation. Surprisingly, this signal transduction pathway also suppressed the premature formation of GCs by inhibiting spontaneous B cell activation (Walsh EMBO Reports 2015). In sum, we uncovered and continue to study a signal transduction pathway and genetic program that controls the initiation, progression, and termination of an effective humoral immune response (Waters Cell Cycle 2015). Errors in this signal transduction pathway or genetic program provide known and novel candidates for uncontrolled B cell growth, survival, and malignant transformation.
Current lab project areas include:
(Techniques include ChIP, next-generation sequencing, epigenetic approaches, IP-mass spectrometry, mouse models, quantitative imaging, and many standard cellular and molecular biology approaches)
(1) Dissection of CRTC2 regulation of B lymphocyte activation, differentiation, and function
(2) Evaluation of a mouse model that is defective for antigen-driven plasma cell differentiation
(3) Molecular elucidation of the role for LKB1 and AMPK in starting GC reactions
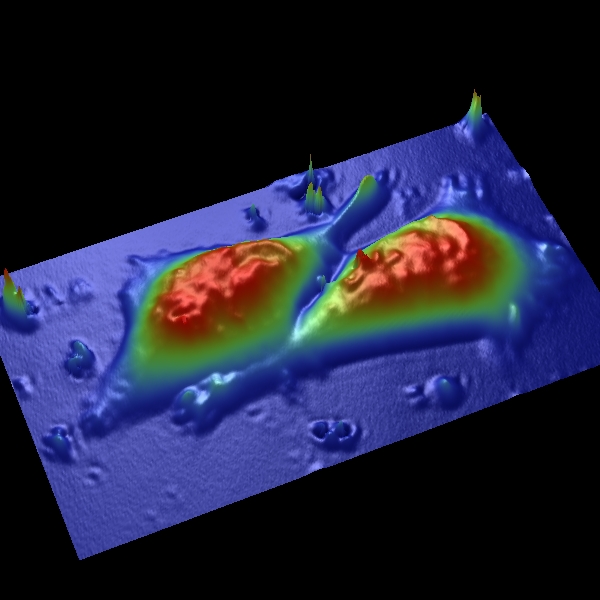
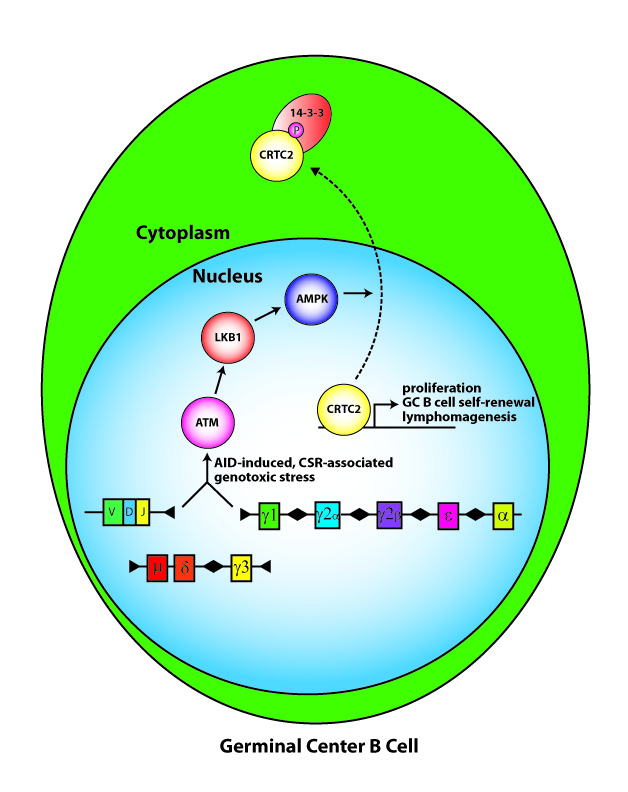
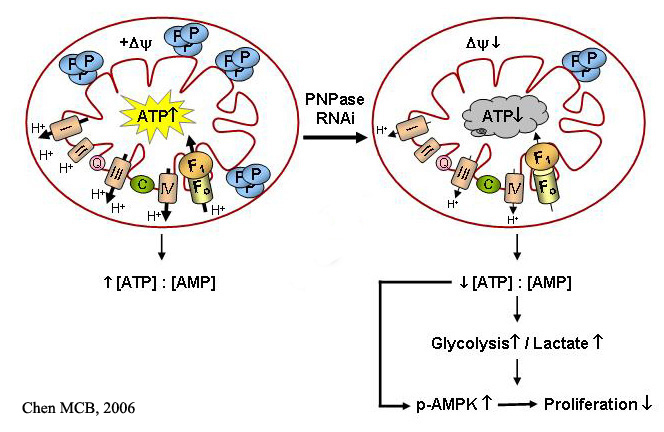

A new area of investigation links developing expertise in the group in two areas- mitochondrial and human embryonic stem cell (hESC) biology. The rationale for these studies is the need to understand how changes in metabolism, which normally shifts from glycolysis to aerobic respiration as fertilized ova implant during embryogenesis, shifts back again to a main dependence on glycolysis (the Warburg effect) with malignant transformation. We reason that dynamic changes in mitochondria function and hESC dependence on oxygen tension will mimic key features of malignant degeneration. Current studies include evaluating the role of native and damaged mitochondria in normoxic and hypoxic oxygen tension environments to determine a role in controlling hESC decisions for self-renewal, differentiation, or quiescence, which could resemble similar decisions in “cancer stem cells”.